通专融合AGI

具备一百万词元(Token)的超长文本窗口及开源模型中领先的推理能力,并支持自主规划和在线信息整合的开源大语言模型。

具备一百万词元(Token)的超长文本窗口及开源模型中领先的推理能力,并支持自主规划和在线信息整合的开源大语言模型。
十亿~千亿
模型参数
5T
模预训练语料 (tokens)
1M
语境窗口长度


支持图像、视频、文字、语音、三维点云等模态处理的开源多模态大模型,具备强大的真实世界感知能力,首创的渐进式对齐训练策略,实现了成本更低、性能更高。

支持图像、视频、文字、语音、三维点云等模态处理的开源多模态大模型,具备强大的真实世界感知能力,首创的渐进式对齐训练策略,实现了成本更低、性能更高。
10亿~780亿
开源参数
MMMU突破70%
强劲 全面 普惠


具备长思维能力,并能在推理过程中进行自我反思和纠正,从而在数学、代码、推理谜题等多种复杂推理任务上取得更优结果。

具备长思维能力,并能在推理过程中进行自我反思和纠正,从而在数学、代码、推理谜题等多种复杂推理任务上取得更优结果。
自主生成高智力密度数据
具备元动作思考能力
采用链式推理策略
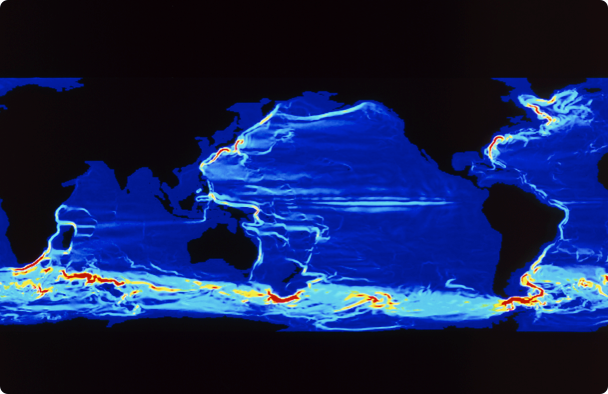


全球首个人工智能驱动的全尺度、全要素的气象海洋预报大模型体系,包含短临、中期、多年-年际尺度等预报模型,覆盖海陆空多种核心要素。

全球首个人工智能驱动的全尺度、全要素的气象海洋预报大模型体系,包含短临、中期、多年-年际尺度等预报模型,覆盖海陆空多种核心要素。
中期预报有效时效首次突破10天
推动全球气象预报发展提速10倍
大模型预报分辨率首次进入10千米


AI谱学大模型,助力研究者更高效、更精准地探索微观世界,推动物质科学研究创新。

AI谱学大模型,助力研究者更高效、更精准地探索微观世界,推动物质科学研究创新。
AI+EPR
秒级解谱、识谱寻源


具备强大理解和分析能力的种业大模型,降低了种领域的学习门槛,助力提升从业者的研究、实践效率。

具备强大理解和分析能力的种业大模型,降低了种领域的学习门槛,助力提升从业者的研究、实践效率。
结合书生·浦语2.0
生物育种垂直领域
品种选育、农艺性状、推广区域、栽培技术


全面由国产新兴算力异构互联技术支撑,完整训练而出的全能多模态生成大模型。

全面由国产新兴算力异构互联技术支撑,完整训练而出的全能多模态生成大模型。
异构混训
效率达97.5%
提升1.6倍
虚实结合物理智能


仅通过单一平台、简单代码输入,实现仿真环境中机器人灵活训练,提升数据采集效率,为研究者提供“一站式”具身智能开发解决方案。

仅通过单一平台、简单代码输入,实现仿真环境中机器人灵活训练,提升数据采集效率,为研究者提供“一站式”具身智能开发解决方案。
89种功能性场景
无需协作数据,先易后难学习策略
十万可交互数据高精视觉仿真,物理驱动可交互,高效并行训练

生成式世界模型AETHER全部由合成数据训练而成,不仅在传统重建与生成任务中表现领先,更首次赋予大模型在真实世界中的 3D 空间决策与规划能力,可助力机器人完成目标导向的视觉规划、4D 动态重建、动作条件的视频预测等复杂任务。
生成式世界模型AETHER全部由合成数据训练而成,不仅在传统重建与生成任务中表现领先,更首次赋予大模型在真实世界中的 3D 空间决策与规划能力,可助力机器人完成目标导向的视觉规划、4D 动态重建、动作条件的视频预测等复杂任务。
目标导向视觉规划
4D 动态重建
动作条件视频预测
安全可信AI
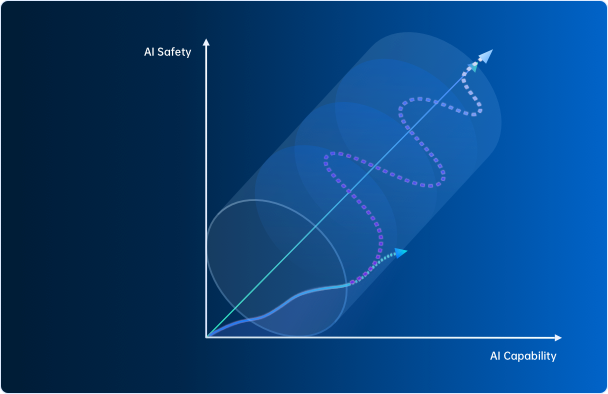
AI-45°平衡律
上海人工智能实验室主任、首席科学家,清华大学惠妍讲席教授周伯文在会上提出以“AI-45°平衡律” (AI-45° Law)这一技术体系,实现兼顾安全与性能的可信AGI。
AI-45°平衡律
上海人工智能实验室主任、首席科学家,清华大学惠妍讲席教授周伯文在会上提出以“AI-45°平衡律” (AI-45° Law)这一技术体系,实现兼顾安全与性能的可信AGI。
平衡
指短期可以有波动,但不能长期低于45°(如同现在),也不能长期高于45°(这将阻碍发展与产业应用)
要求
强技术驱动、全流程优化、多主体参与以及敏捷治理
发展
泛对齐→可干预→能反思 三个递进阶段
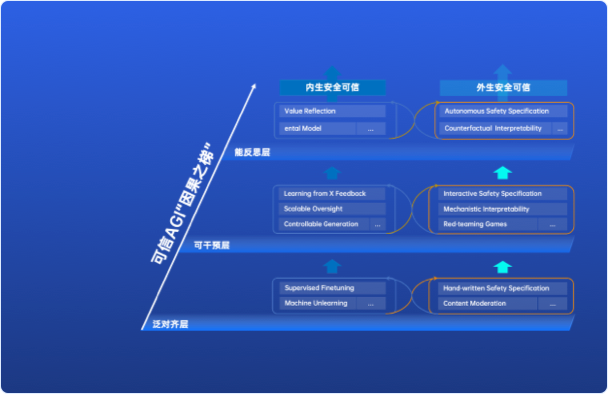
因果之梯
为了探索AI-45°平衡律的实际应用,上海AI实验室提出了基于“因果阶梯”的三层技术框架,旨在解决通往通用人工智能(AGI)过程中关于安全性和可信度的关键要求。
因果之梯
为了探索AI-45°平衡律的实际应用,上海AI实验室提出了基于“因果阶梯”的三层技术框架,旨在解决通往通用人工智能(AGI)过程中关于安全性和可信度的关键要求。
核心目标
促进AGI系统的渐进式发展,从泛对齐和干预能力演变到自我反思,从而确保较高的安全性和可信度
关键维度
内生安全可信、外生安全可信
评测体系


面向大模型的开源方和使用者, 提供开源、高效、全面的大模型评测开放平台。 为大语言模型、多模态模型等各类模型提供一站式评测服务。全面量化模型在知识、语言、理解、推理和考试等五大能力维度的表现,客观中立地为大模型技术创新提供坚实的技术支撑。

面向大模型的开源方和使用者, 提供开源、高效、全面的大模型评测开放平台。 为大语言模型、多模态模型等各类模型提供一站式评测服务。全面量化模型在知识、语言、理解、推理和考试等五大能力维度的表现,客观中立地为大模型技术创新提供坚实的技术支撑。
CompassHub
评测集社区:提供高时效性,高质量评测集
CompassRank
性能榜单:发布权威榜单,洞悉行业趋势
CompassKit
评测工具:支撑高效评测,支持能力分析

医疗大模型应用检测验证中心
全国首个医疗大模型应用检测验证中心,打通 “训、评、用一体化”链路,为产业应用提供方向牵引与安全保障,促进生态发展。检测验证中心已完成上海12家医院的医疗大模型应用场景评测,规范产业和促进产业升级。
医疗大模型应用检测验证中心
全国首个医疗大模型应用检测验证中心,打通 “训、评、用一体化”链路,为产业应用提供方向牵引与安全保障,促进生态发展。检测验证中心已完成上海12家医院的医疗大模型应用场景评测,规范产业和促进产业升级。
覆盖国家《卫生健康行业人工智能应用场景参考指引》指导文件中32个场景辐射的10多个亚场景应用评测
基础平台
行业应用
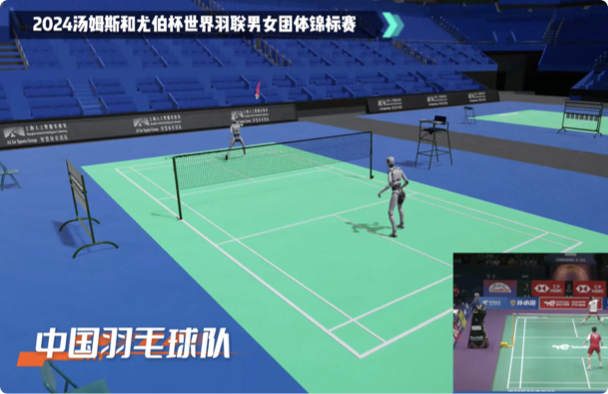

国内首个AI体育技战术大模型,仅基于体育运动的视频画面,即可实时获取并分析出关键参数信息,为体育比赛直播及运动员竞技水平分析提供AI支持。

国内首个AI体育技战术大模型,仅基于体育运动的视频画面,即可实时获取并分析出关键参数信息,为体育比赛直播及运动员竞技水平分析提供AI支持。
仅基于单路电视PGM信号(最终播出信号)即可获取技战术指标
动态时间切片技术(Dynamic Figure Out)


基于央视听媒体大模型(MediaGPT),与中央广播电视总台联合制作首部AIGC系列动画片《千秋诗颂》,国内观众超1亿,触达海外观众11亿,英、西、法、阿等10余个语种版本,同步在全球70余家主流媒体播出。

基于央视听媒体大模型(MediaGPT),与中央广播电视总台联合制作首部AIGC系列动画片《千秋诗颂》,国内观众超1亿,触达海外观众11亿,英、西、法、阿等10余个语种版本,同步在全球70余家主流媒体播出。


















